 AJAZ UL H WANI
AJAZ UL H WANI
Principal Investigator/Scientist-D
Wellcome Trust-DBT Intermediate Fellow,
Ramanujan Fellow
ahwani@kashmiruniversity.ac.in
Ph. D
National Centre for Biological Sciences-TIFR (NCBS-TIFR), Bangalore
Postdoctoral Fellow
Harvard University, USA
(2009-2014)
Research
Eukaryotic genomes are packaged as chromatin within the cell nucleus. Chromatin has a multiscale three-dimensional architecture varying from nucleosomes, loops, topologically associating domains to chromosome territories. The three-dimensional organization of chromatin within the nuclear space has been linked with genome function like regulation of gene expression and other DNA associated processes. Although application of various methods continue to provide fine details of chromatin architecture but the principles governing the three-dimensional organization of chromatin are just beginning to unravel. We are interested in understanding the role of chromatin-associated proteins in regulating chromatin organization at different genomic and special scales, from nucleosome positioning/accessibility to chromatin topology. To address this problem we are using an interdisciplinary approach including protein biochemistry/biophysics, genome wide sequencing based methods (like ChIP-seq, Chromosome conformation capture, MNase-seq and RNA-seq), mass spectrometry to polymer modeling.
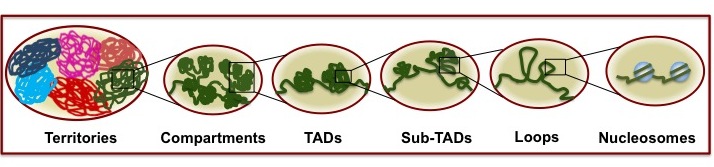
Hierarchical organization of genome (Shah et al.,2018 Nucleus (Taylor and Francis))
The hierarchical model of genome organization is the most accepted model in field. In a hierarchical organization, different levels have dependences on each other and alterations in one level should get transmitted toother levels lying below or above it, resulting in "Bottom-Up" and "Top-Down" causation properties. We are testing the strength of the hierarchical nature of 3D chromatin orgnization by perturbing a particular level and monitor its effects on orther levels of organization. Our data show that disruption of self-association of a polycomb protein, polyhomeotic, PH (PH-PH interaction) disrupts chromatin contacts, decreases nucleosome occupancy and alter accessibility. Simulations investigating the interplay between distant chromatin contacts and nucleosome occupancy, both of which are regulated by PH polymerization, suggest that nucleosome density increases when contacts between different regions of chromatin are established. These results suggest that higher-order organization (chromatin contacts) modulate lower level of organization, nucleosomes -"Top-Down" causation. (Amin et al., Life-Science Alliance, In Press)
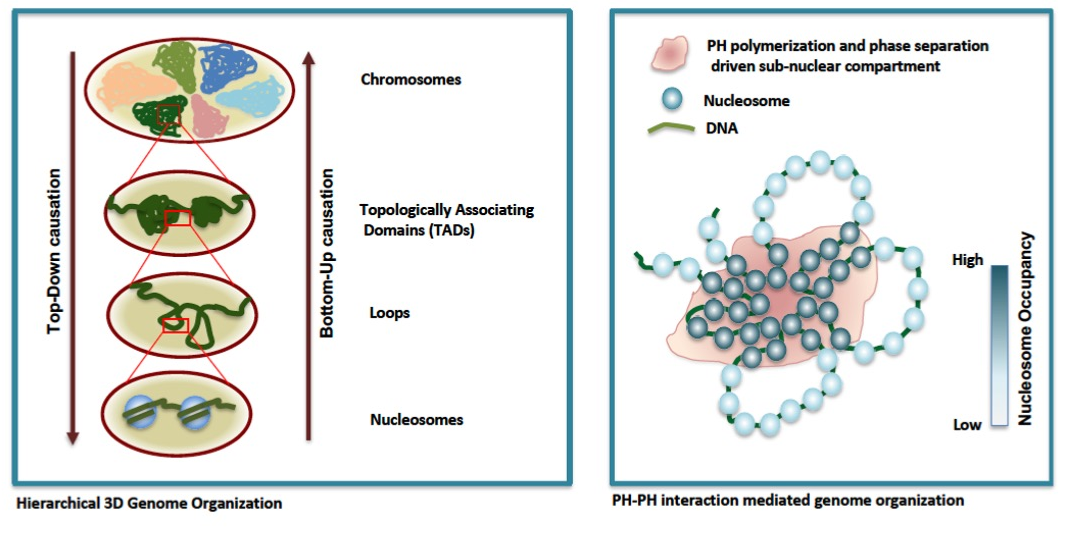
Group Members:
Adfar Amin (Ph. D student)
Suhail Ahmad Shiekh (Ph. D student)
Abas Ahmad Dar (Ph. D student)
Raj Wamsik (M. Sc. student)
Wardha Shah (M. Sc. student)
Humaira (M. Sc. student)
Publications:
1. Fouziya, S.1†, Krietenstein, N.2†, Mir, U. S.,3† Mieczkowski, J.4, Khan, M. A1, Baba, A.1, Dar, M. A.1, Altaf, M.3 & Wani A. H.1* Genome wide nucleosome landscape shapes 3D chromatin organization. Science Advances 2024 (In press)
,
2. Younus A Bhat, Javaid Y Bhat,Shajrul Amin, Jayant B Udgaonkar & Wani A. H.* HDX-MS reveals conformational dynamics of chromatin remodeler ISWI in resting state and during nucleosome sliding (biorxiv, doi.org/10.1101/2023.07.30.549839
3. Amin, A.†, Kadam, S†, Mieczkowski, J., Ahmad, I., Bhat, Y. A., Shah, F., Tolstorukov, M., Kingston, R. E., Padinhateeri, R. & Wani A. H.* Disruption of Polyhomeotic polymerization decreases nucleosome occupancy and alters genome accessibility Life-Science Alliance 2023 Vol.6, e202201768
4. Michieletto, D., Marrenduzzo, D. & Wani A. H.* Chromosome-wide simulations uncover folding pathway and 3D organization of interphase chromosome (Bioarchives)
5. Shah, F. R., Bhat, Y. A. & Wani, A. H.* Subnuclear distribution of proteins: Links with genome architecture. Nucleus, 2018 Jan 1;9(1):42-55. doi: 10.1080/19491034.2017
6.Wani, A. H., Boettiger, A. N., Shorderet, P., Ergun, A., Munger, C., Saderyev, R. I., Zhuang, X., Kingston, R. E. & Francis, N. J. * Chromatin topology is coupled to Polycomb Group protein subnuclear organization. Nature Commu., 2016 7:10291doi: 10.1038/ncomms10291.
7. Follmer. N. E., Wani, A. H., & Francis, N. J. *. Polycomb group proteins remain bound at specific sites during mitosis. PloS Genetics. 2012, 8,12: e1003135.
8. Wani, A. H. and Udgaonkar, J. B.* Mass spectrometry studies of protein folding. Current Science, 2012, 102, 2, 245-265.
9. Wani, A. H. and Udgaonkar, J. B. * Native state dynamics drive the unfolding of the SH3 domain of PI3 kinase at high denaturant concentration. Proc Natl Acad Sci USA. 2009, 106, 49: 20711-20716.
10. Wani, A. H. and Udgaonkar, J. B. * Revealing a concealed intermediate that forms after the rate-limiting step of refolding of the SH3 domain of PI3 kinase. Journal of Molecular Biology, 2009, 387, 2: 348-362.
11. Wani, A. H. and Udgaonkar, J. B. * HX-ESI-MS and optical studies of the unfolding of thioredoxin indicate stabilization of a partially unfolded, aggregation-competent intermediate at low pH. Biochemistry, 2006,45, 37, 11226-11238
12. Modi, S., Wani, A. H. & Krishnan Y. * The PNA-DNA hybrid I-motif: implications for sugar-sugar contacts in i-motif tetramerization. Nucleic acids research, 2006, 34, 16, 4354-4363.
Teaching
1. Systems Biology: Netwroks and Noise
2. Spectroscopy and Microscopy
3. Biochemical Techniques
Awards/Fellowships:
- Awarded Wellcome Trust-DBT intermediate fellowship by Wellcome Trust-DBT India Alliance. (2016)
- Awarded Early Career Research Award by Department of Science and Technology, India. (2016)
- Awarded Ramanujan fellowship by Department of Science and Technology, India. (2014)
Alumini:
Postdocs:
Zaffer Ullah Zargar (Presently: Assistant Professor High Education Deptt. J&K)
Ph D Students:
Younus Ahamd Bhat (Presently: Postdoctoral Fellow, University of Massachuesetts, USA, (Prof Craig Peterson lab.))
Fouziya Shah
Masters Students:
Masood Ahmad Khan
Ateeb Shafi
Mehvish Rafiqui
Aqsa Ellahi
Aemon Baba
Wajahat Waseem
Mumtaz Arif
Safiya Majeed
Iqra Jan
Funding

New layer...
New layer...